Research Theme
2.Development of non-target lipidomics platform
Kazutaka Ikeda, Makoto Arita
Laboratory for Metabolomics, RIKEN Center for Integrative Medical Sciences
Lipid metabolites consist of a wide variety of fatty acids or of polar groups, and some of them are quantitatively and qualitatively variable or oxidizable depending on the situation. To investigate lipid metabolic changes in detail, lipidomics analysis by LC-MS has been wildly applied. However, it remains insufficient to measure them globally and difficult to discover the important metabolic change unbiasedly by conventional approach.
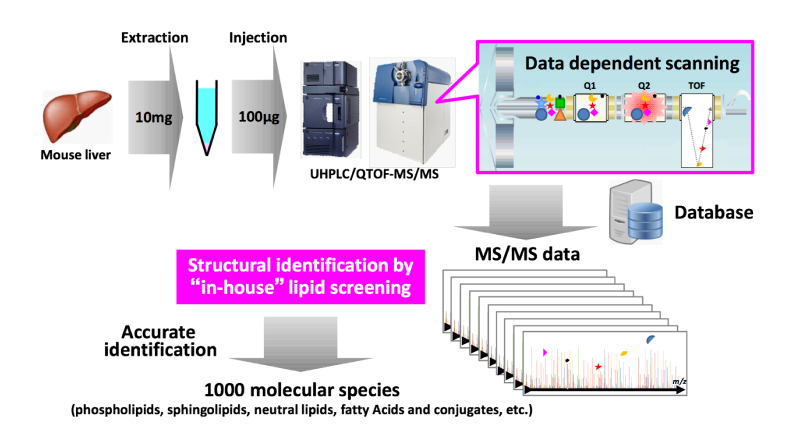
To solve these problems, we constructed a new lipidomics platform by quadrupole time-of flight (Q-TOF) mass spectrometer which provided the ability to perform non-target analyses with high resolution and accurate MS/MS information. MS/MS trigger was set at low threshold level in order to detect as globally as possible, and lipid structures are unbiasedly identified by in-house lipid database screening. Conventional screening techniques by using “in silico” MS/MS spectra library have a problem with the low identification accuracy. In our screening system, MS/MS fragment peaks necessary for accurate identification were carefully selected by using a measured MS/MS spectra library and optimally weighted to eliminate misidentification. For example, mouse liver extract (equivalent to 100 μg) was injected into our LC-MS system and about 1000 molecular species (phospholipids, sphingolipids, neutral lipids, fatty acids and conjugates, etc.) were accurately and efficiently detected by our database screening.
Non-target lipidomics platform enables us to discover potential link between lipid metabolism and biological phenotypes. Our new approach has a strong potential to search for lipids of interest globally, or to identify unknown lipid species in non-biased fashion.